13. Modeling Volumes and Multiple Compartments
This notebook example provides a basic demonstration on how to create and dynamically simulate multi-compartment models.
Illustrated below is the multi-compartment model utilized in this notebook:
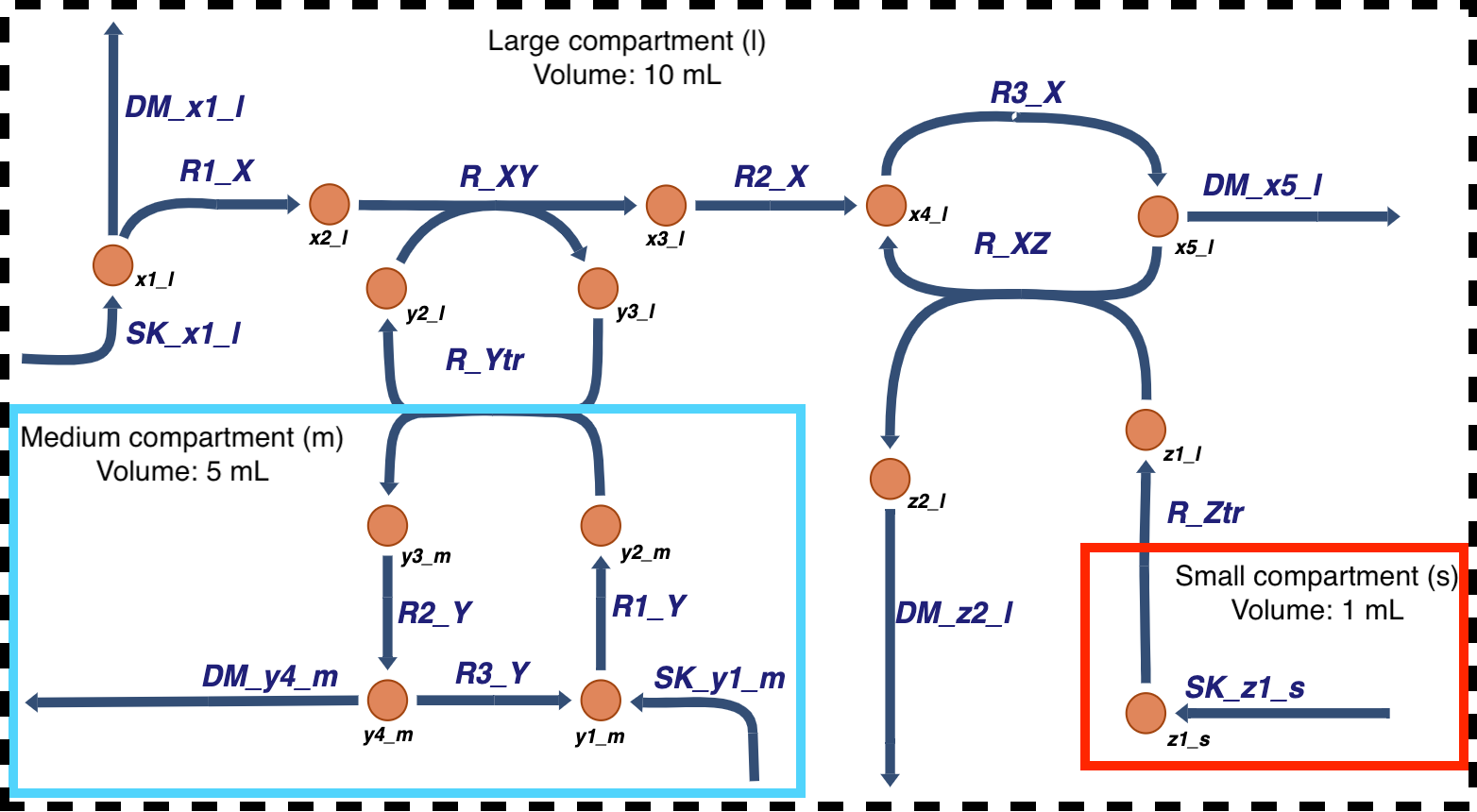
In the above example: * For metabolite x: * The biochemical pathway for the conversion of x occurs in the large compartment outlined by the dotted black line. * For metabolite y: * Cofactor y is necessary for the conversion of metabolite x in the biochemical pathway. * The synthesis of y occurs in the medium compartment outlined by the blue line. * R_Ytr is an antiporter, coupling the import of y2 into the large compartment with the export of
y3 to the medium compartment. * For metabolite z: * Protein z is synthesized in the small compartment outlined by the red line. * Protein z is also to facilitate the conversion of x5 back into x4 and for metabolic functions outside of the model’s scope.
The reaction converting x5 back to x4 converted into x5 The pair of irreversible reactions R3_X and R_XZ form a cycle that is used to The synthesis and degradation of metabolite y occurs in the medium compartment outlined in blue.
COBRApy is currently in the process of developing improved compartment handling. These changes are outlined in the following COBRApy issues:
MASSpy is awaiting these COBRApy changes in order to improve how compartments are handled in dynamic simulations, SBML compatibity, etc. Once these changes have been implemented in COBRApy, a new version of MASSpy will be developed and released with improved functionality around compartments and their handling.
13.1. Models with Multiple Compartments
[1]:
import sympy as sym
from mass import (
MassConfiguration, MassMetabolite, MassModel, MassReaction, Simulation)
from mass.example_data import create_example_model
from mass.visualization import plot_time_profile
model = create_example_model("MultiCompartment")
Set parameter Username
13.1.1. Viewing compartments in a model
The MassModel.compartments attribute is used to get dict with compartment identifiers and their corresponding names.
[2]:
model.compartments
[2]:
{'l': 'Large', 'm': 'Medium', 's': 'Small'}
The names for the compartments can be reset or changed by using the MassModel.compartments attribute setter method. To reset compartment names, pass an empty dict:
[3]:
model.compartments = {}
To set a new name for a compartment, set a dict using the MassModel.compartments method with the compartment identifer as the key and the compartment name as the value. Compartments can be set one at a time, or multiple at once:
[4]:
model.compartments = {"l": "the large compartment"}
print(model.compartments)
model.compartments = {"m": "the medium compartment", "s": "the small compartment"}
print(model.compartments)
{'l': 'the large compartment', 'm': '', 's': ''}
{'l': 'the large compartment', 'm': 'the medium compartment', 's': 'the small compartment'}
13.1.1.1. Volume units
To get a list of all UnitDefinition(s) that contain a volume base unit, an modified filter that scans the base units can be applied:
[5]:
def volumes_filter(udef):
if list(filter(lambda u: u.kind in ["liter","litre"], udef.list_of_units)):
return True
return False
print(model.units.query(volumes_filter))
[<UnitDefinition Milliliters "mL" at 0x7fd600f0c190>, <UnitDefinition Concentration "mol_per_mL" at 0x7fd600f0c2b0>]
13.1.2. Enabling compartment volumes in rate laws
By default, the MassConfiguration.exclude_compartment_volumes_in_rates is set as True.
[6]:
mass_config = MassConfiguration()
print(mass_config.exclude_compartment_volumes_in_rates)
True
Therefore, all automatically generated mass action rate laws do not include the compartment volume:
[7]:
print(model.reactions.get_by_id("R2_X").rate)
kf_R2_X*x3_l(t)
To enable compartment volumes in rate laws, the MassConfiguration.exclude_compartment_volumes_in_rates attribute must be set to False.
[8]:
mass_config.exclude_compartment_volumes_in_rates = False
print(model.reactions.get_by_id("R2_X").rate)
kf_R2_X*volume_l*x3_l(t)
As seen above, volume parameters are added into the rate laws to represent compartment volumes. The volume parameters have identifiers of format volume_CID , with CID referring to the compartment identifier (e.g., “l” for large compartment). For a reaction that crosses compartments, more than one “volume” parameter will appear as a variable in the rate:
[9]:
for param in model.reactions.get_by_id("R_Ytr").rate.atoms(sym.Symbol):
if str(param).find("volume") != -1:
print(param)
volume_m
volume_l
See the section on Excluding compartments from rates in the Global Configuration tutorial for more information about the exclude_compartment_volumes_in_rates attribute.
13.1.3. The “boundary” compartment
In boundary reactions (e.g., pseudeoreactions such as sinks, demands, and exchanges), metabolites that exist in the boundary a.k,a. the boundary conditions, are given a default “boundary” compartment with the identifier “b”. This compartment is treated as a pseudo-compartment, and therefore the ‘boundary’ metabolites are treated as pseudo-metabolites, meaning no corresponding object is created for them.
Boundary metabolites can be accessed either through the MassReaction.boundary_metabolite method.
[10]:
x1_b = model.reactions.get_by_id("SK_x1_l").boundary_metabolite
x1_b
[10]:
'x1_b'
If a reaction is not a boundary reaction (i.e., MassReaction.boundary==False) then None will be returned:
[11]:
print(model.reactions.get_by_id("R_Ytr").boundary_metabolite)
None
The boundary_metabolite attribute is useful for getting and setting values in the MassModel.boundary_conditions attribute.
[12]:
model.boundary_conditions[x1_b] = 2
model.boundary_conditions
[12]:
{'x1_b': 2}
To change the ‘boundary’ compartment identifier and name, a dict is passed to the MassConfiguration.boundary_compartment attribute setter:
[13]:
print("Before: {0}\n{1}".format(mass_config.boundary_compartment, model.boundary_metabolites))
mass_config.boundary_compartment = {"xt": "External compartment"}
print("\nAfter: {0}\n{1}".format(mass_config.boundary_compartment, model.boundary_metabolites))
Before: {'b': 'boundary'}
['x1_b', 'x5_b', 'y1_b', 'y4_b', 'z1_b', 'z2_b']
After: {'xt': 'External compartment'}
['x1_xt', 'x5_xt', 'y1_xt', 'y4_xt', 'z1_xt', 'z2_xt']
The “boundary” compartment is automatically assumed to have a volume of 1, and therefore is not factored in the rate laws. It is also ignored by the MassModel.compartments attribute, even when explicitly set:
[14]:
for r in model.sinks:
print("{0}: {1}".format(r.id, r.get_mass_action_rate()))
model.compartments = {"xt": "External compartment"}
model.compartments
SK_y1_m: kf_SK_y1_m*y1_xt
SK_z1_s: kf_SK_z1_s*z1_xt
[14]:
{'l': 'the large compartment',
'm': 'the medium compartment',
's': 'the small compartment'}
See the section on For compartments and SBML in the Global Configuration tutorial for more information about the boundary_compartment attribute.
The ‘boundary’ pseudo-compartment and ‘boundary’ pseudo-metabolites are designed to make working with boundary conditions convenient at the cost of having finer user control. This primarily useful for * Setting functions as boundary conditions (e.g., an oscillating function for external oxygen concentration) * Using custom rates to set fixed inputs, causing irrelevant boundary conditions to be ignored altogether.
However, for finer control over external compartment and boundary conditions (and general best practices for SBML compatibility in MASSpy), it is recommended to (1) create new MassMetabolite objects, define their compartment and initial_condition attributes, (2) set the fixed attribute as True, and (3) add the metabolites to the appropriate reactions. This ensures the concentration of the metabolite is fixed at a constant value, and that its initial condition value is treated
as a boundary condition.
13.1.3.1. Fixed inputs
To bypass using the ‘boundary’ pseudo-compartment, it is recommended to set a fixed input using a custom rate law:
[15]:
for r in model.reactions.get_by_any(["SK_x1_l", "SK_y1_m", "SK_z1_s"]):
model.add_custom_rate(r, custom_rate=r.kf_str)
print("{0}: {1}".format(r.id, r.rate))
SK_x1_l: kf_SK_x1_l
SK_y1_m: kf_SK_y1_m
SK_z1_s: kf_SK_z1_s
13.2. Getting and setting compartment volumes
Support for compartment volumes is currently through the MassModel.custom_parameters attribute. To view what compartment volumes are set:
[16]:
def volume_filter(parameter):
if str(parameter).startswith("volume"):
return True
return False
for vol_id in filter(volume_filter, model.custom_parameters):
print("{0}: {1}".format(vol_id, model.custom_parameters[vol_id]))
volume_l: 10.0
volume_m: 5.0
volume_s: 1.0
To set or change a compartment volume, the value in the MassModel.custom_parameters dict is set using the volume parameter ID as the key:
[17]:
# Set the large compartment volume to 15
model.custom_parameters["volume_l"] = 15
# Double current medium compartment volume
model.custom_parameters["volume_m"] = model.custom_parameters["volume_m"] * 2
# 10% decrease to current small compartment volume
model.custom_parameters["volume_s"] = model.custom_parameters["volume_s"] * (1 + (-10/100))
for vol_id in filter(volume_filter, model.custom_parameters):
print("{0}: {1}".format(vol_id, model.custom_parameters[vol_id]))
volume_l: 15
volume_m: 10.0
volume_s: 0.9
13.3. Simulating with Volumes and Multiple Compartments
Using a newly loaded model, the following section provides guidance on dynamic simulations for models with multiple compartments and includes examples on perturbing compartment volume.
[18]:
# Ensure compartments are active and boundary compartment is reset
mass_config.exclude_compartment_volumes_in_rates = False
mass_config.boundary_compartment = {'b': 'boundary'}
# Start with a fresh model, checking to ensure compartment volumes are reset
model = create_example_model("MultiCompartment")
for vol_id in filter(volume_filter, model.custom_parameters):
print("{0}: {1}".format(vol_id, model.custom_parameters[vol_id]))
volume_l: 10.0
volume_m: 5.0
volume_s: 1.0
As always, a model must first must be loaded into the mass.Simulation. object in order to run a simulation. A quick simulation shows that the model is already at a steady state:
[19]:
simulation = Simulation(model, verbose=True)
conc_sol = simulation.simulate(model, time=(0, 1000))[0]
plot_time_profile(conc_sol, plot_function="loglog", legend="right outside")
Successfully loaded MassModel 'MultiCompartment' into RoadRunner.
[19]:
<AxesSubplot: >
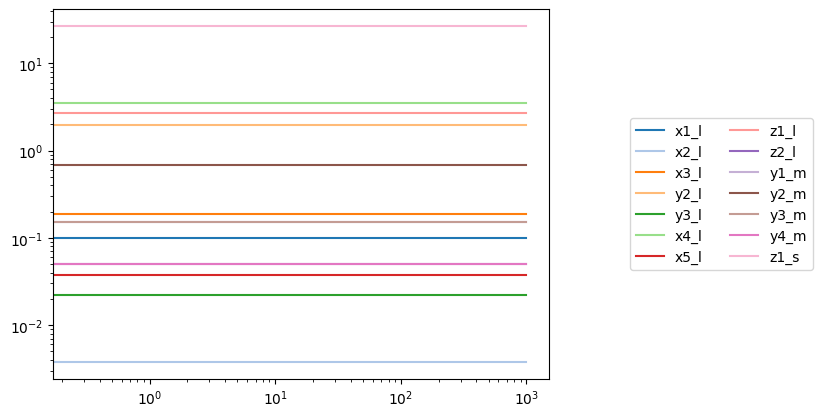
A volume parameter can be perturbed just like any other parameter using a dict. For example, suppose volume of the large compartment volume_m lost 40% of its volume:
[20]:
conc_sol = simulation.simulate(model, time=(0, 1000), perturbations={
"volume_m": "volume_m * (1 - 0.4)",
})[0]
plot_time_profile(conc_sol, plot_function="loglog", legend="right outside")
[20]:
<AxesSubplot: >
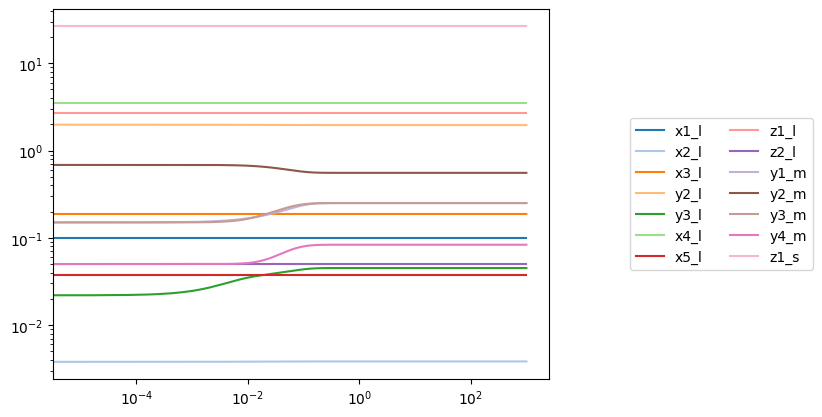
Note that in the above simulation, several of the metabolite concentrations in the large compartment changed. The observable argument can be used with the MassMetabolite.compartment attribute to look at metabolites in a specific compartment for futher examination
[21]:
plot_time_profile(conc_sol, observable=list(model.metabolites.query(lambda m: m.compartment == "m")),
plot_function="semilogx", legend="right outside")
[21]:
<AxesSubplot: >
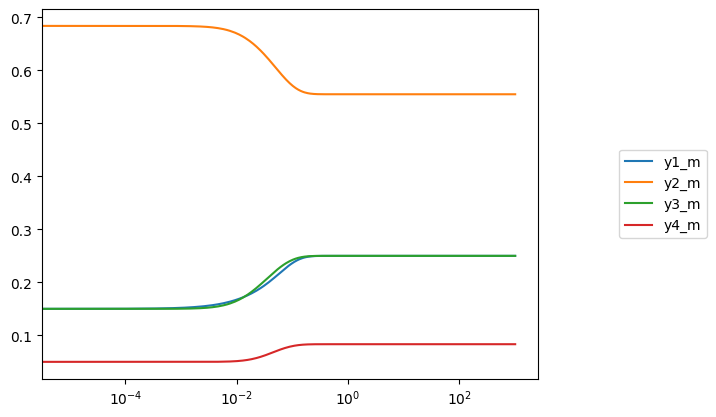
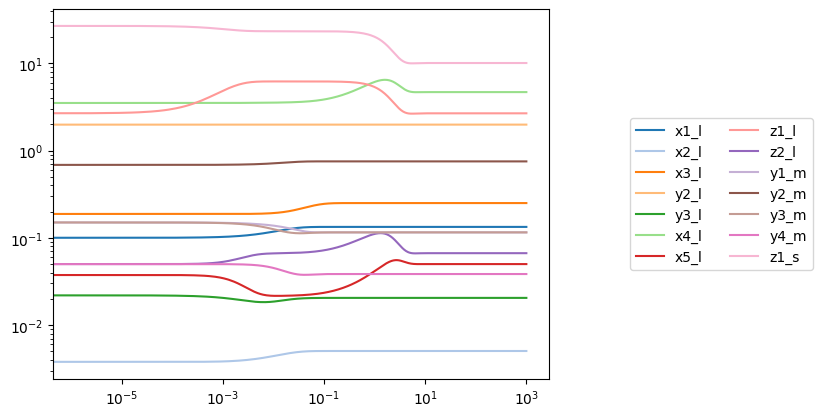
Multiple volumes also can be perturbed simultaneously. For example, suppose 1 mL of fluid from the large compartment was transfered to the small compartment, while 1.5 mL was transfered to the medium compartment:
[22]:
conc_sol = simulation.simulate(model, time=(0, 1000), perturbations={
"volume_l": "volume_l - 2.5",
"volume_m": "volume_m + 1.5",
"volume_s": "volume_s + 1.0"
})[0]
plot_time_profile(conc_sol, plot_function="loglog", legend="right outside")
[22]:
<AxesSubplot: >
Helpful tips: When enabling compartment volumes, it is up to the user to track their units to ensure that no numerical consistency issues arise. To make this a bit easier, be aware of the following MASSpy expectations and behaviors:
When compartment volumes are disabled, MASSpy expects that volumes are already factored into initial condition values, and therefore considers values to be initial concentrations. Consequently, metabolite solutions returned by solutions will be for metabolite concentrations (e.g., mol/L, g/cDW)
When compartment volumes are enabled, MASSpy expects that volumes have not been factored factored into initial condition values, and therefore considers values to be initial amounts. Consequently, metabolite solutions returned by solutions will be for metabolite amounts (e.g., mol, grams)